It is easy to download bioinformatics resources into Bioclipse using the
WSDbfetch Web service at the
European Bioinformatics Institute (EBI). From console you can use the method:
ws.downloadDbEntry(String db, String query, String format)From the GUI you can use menu:
File > New... and select the
Wizard Download > Query WSDbfetch at EBI. Click next and see the many available databases in the dropdown list Supported databases:
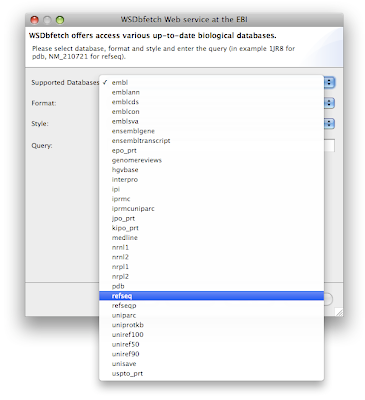
Try for example to download the sequence "NM_210721" from the
"refseq" database.
If you'd like to download proteins in
PDB format, there is a convenience wizard available in the New Wizards dialog available from the menu
File > New...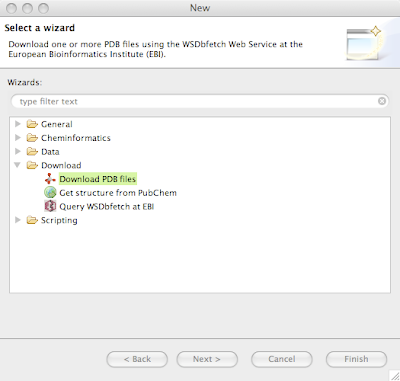
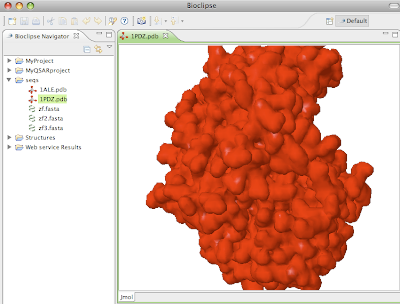
Clicking
Next allows for inputting a comma-separated list of PDB IDs, try for example "1ale,2pdz". Clicking next downloads the file to the selected folder in the Navigator, or to the Virtual project if nothing is selected. Below is visualized 2PDZ in the Jmol Editor.