2012-12-22
Bioclipse 2.6 released
2012-12-13
Bioclipse 2.6.0 Release Candidate is out
The Bioclipse team is proud to announce a release candidate for Bioclipse with version 2.6.0.RC1.
We kindly ask from the community to put Bioclipse to the test and report bugs at http://bugs.bioclipse.net.
Download Bioclipse 2.6.0.RC1.
A getting-started guide is available.
Summarizing what's new (in no particular order):
- New Decision Support models (e.g. chemspider, daphnia, herg, opentox, SmartCyp)
- Bioclipse-R, including R-models for DS
- New online updates/installation
- New user interface for downloading data and predictive models
- Installer for Windows and Mac OS X
- CDK version 1.4.15 including e.g. better double bond localization, more atom types, and speed improvements.
- InChI (jni-inchi 0.8) supporting InChI v 1.0.3 (both standard and non-standard InChI).
- Improved core stability (Eclipse 3.8)
- Nightly build with Hudson and new provisioning system p2
- Numerous bug fixes
2012-04-19
Chemical decision support with Chemspider and ChEMBL-RDF
- Query ChemSpider using the SimilaritySearch via the Web API (SOAP).
- For a maximum of 15 nearest neighbors in Chemspider, query ChEMBL-RDF for interactions, and present information about assays, targets, and interaction values.

2012-04-18
Postdoc position in predictive toxicology at Uppsala University
Postdoc position in cheminformatics, bioinformatics, or computer science - applications in predictive toxicology
We have an open postdoc position in the group of Pharmaceutical Bioinformatics at the Department of Pharmaceutical Biosciences, Uppsala University, Sweden.
The successful applicant will conduct research on data interoperability for predictive toxicology, and especially design and implement an infrastructure consisting of a database and user interfaces for data and predictive models in toxicology. Of particular interest will be to merge chemical and biological data within a semantic framework, and link toxicity data to genomics and metabolomics data (toxicogenomics) with a connection to the Bioclipse framework (www.bioclipse.net). PhD degree or equivalent scholarly competence in a relevant branch of chem/bioinformatics or computer science and a strong interest in informatics and data integration is required. Required competences include web programming, databases, and working knowledge in Java. Experience with linked data is desirable.
Deadline for application: May 9th , 2012.
Link to job ad and application form: http://www.uu.se/jobb/others/annonsvisning?tarContentId=186211&languageId=1
2011-03-08
Bioclipse meets OpenTox
2011-03-07
Presenting at Society of Toxicology 2011

2011-03-04
Web seminar on Bioclipse
2010-08-30
Bioclipse 2.4.1 now on update site
To update go to Help -> Software Updates... and select the features you want to install.
Known problem: The splash screen erroneously shows version number to be 2.4.0 after update. (Technical information can be found at bug: 2084 in our Bugzilla)
2010-07-09
Bioclipse 2.4 released
The Bioclipse team is proud to announce the release of Bioclipse version 2.4. The release contains various new features and bug fixes in cheminformatics and drug discovery, including improved QSAR functionality, site-of-metabolism prediction, semantic web functionality, browsing of large compound collections, editing of chemical structures, and numerous bug fixes.
Bioclipse 2.4 is available for 32 and 64 bit versions of Mac OS X, Linux, and 32 bit version of Windows (Bioclipse for 64 bit Windows is currently unavailable, but will be provided as soon as a native Standard InChI is available for 64 bit Windows).
- Download Bioclipse
- Getting started guide
- Planet Bioclipse integrates blogs related to Bioclipse.
2010-07-06
Bioclipse 2.4.0.RC3 is here
http://pele.farmbio.uu.se/bioclipse-devel/
I have a good feeling about this one. I think Bioclipse 2.4 is really close now.
And as usual, if you download and try the release candidate of course you already know that we love to get bug reports in our Bugzilla. :)
2010-06-01
Bioclipse 2.4.0.RC1 is out
http://pele.farmbio.uu.se/bioclipse-devel/
What is new?
Among the main news are:
- New molecules table.
- We are now using Java 1.6
- Eclipse 3.5.2
- CDK 1.3.5
2010-03-07
Bioclipse is finalist for the Eclipse Community Awards 2010

2010-01-28
Bioclipse 2.2 released
* Cheminformatics, with a pure SWT-based chemical 2D editor (JChemPaint) and a lazy-loading molecules table.
* QSAR, supporting local, REST, and XMPP services
* MetaPrint2D for interactive site-of-metabolism prediction for chemical structures
* StructureDB and VScreen: A chemical database with virtual screening functionality
* The new Decision Support feature with graphical reports using BIRT
* Semantic web features
* Bioinformatics, with the new Sequence Editor and sequence alignments via the Kalign Web service (Experimental)
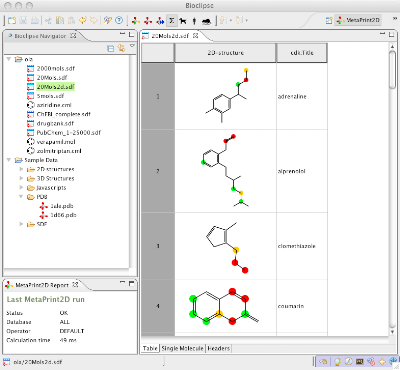 A screenshot from Bioclipse with the MetaPrint2D feature showing predicted sites of metabolsim for a set of drugs in the MoleculesTable.
A screenshot from Bioclipse with the MetaPrint2D feature showing predicted sites of metabolsim for a set of drugs in the MoleculesTable.Note that Bioclipse 2.2.0 requires a fresh download, i.e. it can not be upgraded to by using the software update functionality. A small installation guide is also provided, but the main documentation for Bioclipse is available from help.bioclipse.net; the same information is also available from within Bioclipse from the menu Help > Help Contents. For general questions there is the bioclipse-users and bioclipse.devel mailing lists.
Links:




